Difference between revisions of "Genetic Extinction Technology"
(added quotes) |
m (Text replacement - " -- " to " — ") |
||
| (10 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
{{concept | {{concept | ||
|wikipedia=https://en.wikipedia.org/wiki/Gene_drive | |wikipedia=https://en.wikipedia.org/wiki/Gene_drive | ||
| + | |constitutes=Biological weapon/Research, research, Genetic engineering | ||
|interest_of=Bill & Melinda Gates Foundation, Moderna,Zita Virus,Imperial College London,DARPA,Intelligence Advanced Research Projects Agency,JASON group | |interest_of=Bill & Melinda Gates Foundation, Moderna,Zita Virus,Imperial College London,DARPA,Intelligence Advanced Research Projects Agency,JASON group | ||
}} | }} | ||
| Line 26: | Line 27: | ||
}} | }} | ||
| − | Gene drives can be built from many naturally occurring [[selfish genetic elements]] that use a variety of molecular mechanisms.<ref> | + | Gene drives can be built from many naturally occurring [[selfish genetic elements]] that use a variety of molecular mechanisms.<ref>Champer J, Buchman A, Akbari OS, "Cheating evolution: engineering gene drives to manipulate the fate of wild populations", Nature Reviews. Genetics, volume 17, issue 3, pages 146–59, March 2016, doi = 10.1038/nrg.2015.34</ref> These naturally occurring mechanisms induce similar [[segregation distortion]] in the wild, arising when [[allele]]s evolve molecular mechanisms that give them a transmission chance greater than the normal 50%. |
| − | Most gene drives have been developed in insects, notably mosquitoes, as a way to control insect-borne pathogens. Recent developments designed gene drives directly in viruses, notably [[Herpesviridae|herpesviruses]]. These viral gene drives can propagate a modification into the population of viruses, and aim to reduce the infectivity of the virus.<ref> | + | Most gene drives have been developed in [[insects]], notably [[mosquitoes]], as a way to control insect-borne pathogens. Recent developments designed gene drives directly in viruses, notably [[Herpesviridae|herpesviruses]]. These viral gene drives can propagate a modification into the population of viruses, and aim to reduce the infectivity of the virus.<ref> https://www.nature.com/articles/s41467-020-18678-0</ref><ref>https://www.acsh.org/news/2020/09/30/gene-drives-could-kill-mosquitoes-and-suppress-herpesvirus-infections-15060</ref> |
| − | ==Defense Advanced Research Projects Agency== | + | == History == |
| + | Austin Burt, an [[Extended evolutionary synthesis|evolutionary geneticist]] at [[Imperial College London]], introduced the possibility of conducting gene drives based on natural homing endonuclease [[selfish genetic elements]] in 2003.<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1691325</ref> | ||
| + | |||
| + | Researchers had already shown that such genes could [[Gene-centered view of evolution#Selfish-gene theory|act selfishly]] to spread rapidly over successive generations. Burt suggested that gene drives might be used to prevent a mosquito population from transmitting the [[malaria parasite]] or to crash a mosquito population. Gene drives based on homing endonucleases have been demonstrated in the laboratory in [[transgenic]] populations of mosquitoes<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3093433</ref> and fruit flies.<ref>>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3120159</ref><ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3548849</ref> However, homing endonucleases are sequence-specific. Altering their specificity to target other sequences of interest remains a major challenge.<ref>https://doi.org/10.1038%2Fnrg.2015.34</ref> The possible applications of gene drive remained limited until the discovery of CRISPR and associated RNA-guided endonucleases such as [[Cas9]] and [[CRISPR/Cpf1|Cpf1]]. | ||
| + | |||
| + | In June 2014, the [[World Health Organization]] (WHO) Special Programme for Research and Training in Tropical Diseases<ref>https://www.who.int/tdr/about/en/|access-date=2014-07-18</ref> issued guidelines<ref>https://www.who.int/tdr/news/2014/framwk-eval-gm-mosquitoes/en|access-date=2014-07-18</ref> for evaluating genetically modified mosquitoes. In 2013 the [[European Food Safety Authority]] issued a protocol<ref>http://www.efsa.europa.eu/en/efsajournal/pub/3200</ref> for [[environmental assessment]]s of all [[genetically modified organisms]]. | ||
| + | |||
| + | ==Interested parties== | ||
| + | ===Defense Advanced Research Projects Agency=== | ||
In December [[2017]], documents released under the US [[Freedom of Information Act]] showed that the military research agency [[DARPA]] had invested $100 million in gene drive research,<ref>https://www.theguardian.com/science/2017/dec/04/us-military-agency-invests-100m-in-genetic-extinction-technologies</ref>, making them likely the largest single funder of gene drive research on the planet. The documents also reveal that DARPA either funds or co-ordinates with almost all major players working on gene drive development as well as the key holders of patents on CRISPR gene editing technology. | In December [[2017]], documents released under the US [[Freedom of Information Act]] showed that the military research agency [[DARPA]] had invested $100 million in gene drive research,<ref>https://www.theguardian.com/science/2017/dec/04/us-military-agency-invests-100m-in-genetic-extinction-technologies</ref>, making them likely the largest single funder of gene drive research on the planet. The documents also reveal that DARPA either funds or co-ordinates with almost all major players working on gene drive development as well as the key holders of patents on CRISPR gene editing technology. | ||
{{SMWQ | {{SMWQ | ||
| Line 40: | Line 49: | ||
}} | }} | ||
| − | ==JASON Military Advisory Group== | + | ===JASON Military Advisory Group=== |
The secretive top-level [[JASON group]] of military advisors produced a [[classified]] study on gene drive in [[2017]], reflecting an extremely high level of interest and activity by other sections of the U.S. military and Intelligence community<reF>http://genedrivefiles.synbiowatch.org/2017/12/01/us-military-gene-drive-development/</ref>. | The secretive top-level [[JASON group]] of military advisors produced a [[classified]] study on gene drive in [[2017]], reflecting an extremely high level of interest and activity by other sections of the U.S. military and Intelligence community<reF>http://genedrivefiles.synbiowatch.org/2017/12/01/us-military-gene-drive-development/</ref>. | ||
Emails show that the JASON study was initiated with a two day meeting of a select group of invited gene drive researchers in June 2017. At the meeting, the VP of Global Biotechnology for [[Monsanto]] gave a presentation on crop science and gene drives<ref>http://genedrivefiles.synbiowatch.org/2017/12/01/us-military-gene-drive-development/</ref>. | Emails show that the JASON study was initiated with a two day meeting of a select group of invited gene drive researchers in June 2017. At the meeting, the VP of Global Biotechnology for [[Monsanto]] gave a presentation on crop science and gene drives<ref>http://genedrivefiles.synbiowatch.org/2017/12/01/us-military-gene-drive-development/</ref>. | ||
| − | ==The Intelligence Advanced Research Projects Agency== | + | ===The Intelligence Advanced Research Projects Agency=== |
The [[Intelligence Advanced Research Projects Agency]] (IARPA), an organization within the [[Office of the Director of National Intelligence]], has also expressed interest in funding gene drive work. A scientist involved describes IARPA as “basically the [[intelligence agencies]] version of DARPA, which may be more frightening"<ref>http://genedrivefiles.synbiowatch.org/20170716-fw_-gbird-update-engagements-and-other-important-items-472/</ref> | The [[Intelligence Advanced Research Projects Agency]] (IARPA), an organization within the [[Office of the Director of National Intelligence]], has also expressed interest in funding gene drive work. A scientist involved describes IARPA as “basically the [[intelligence agencies]] version of DARPA, which may be more frightening"<ref>http://genedrivefiles.synbiowatch.org/20170716-fw_-gbird-update-engagements-and-other-important-items-472/</ref> | ||
| − | ==Bill & Melinda Gates Foundation== | + | ===Project for the New American Century=== |
| + | In their 2000 paper: "Rebuilding America's Defences: Strategies, Forces And Resources For A New Century", [[PNAC]] commented on [[biowarfare]] application of a then unnamed technology: | ||
| + | {{SMWQ | ||
| + | |text=And advanced forms of biological warfare that can “target” specific genotypes may transform biological warfare from the realm of terror to a politically useful tool. | ||
| + | |subjects=biological warfare | ||
| + | |authors=Project for the New American Century | ||
| + | |source_details=https://web.archive.org/web/20020923154604/http://www.newamericancentury.org/RebuildingAmericasDefenses.pdf | ||
| + | }} | ||
| + | |||
| + | ===Bill & Melinda Gates Foundation=== | ||
The [[Bill and Melinda Gates Foundation]] paid a PR firm $1.6 million to secretly stack key UN advisory processes with gene drive-friendly scientists, and recruited ostensibly independent academics and public officials into a private collaboration to counteract proposed regulations and to resist calls by scientists and conservationists for an international moratorium<ref>https://etcgroup.org/content/gene-drive-files</ref>. | The [[Bill and Melinda Gates Foundation]] paid a PR firm $1.6 million to secretly stack key UN advisory processes with gene drive-friendly scientists, and recruited ostensibly independent academics and public officials into a private collaboration to counteract proposed regulations and to resist calls by scientists and conservationists for an international moratorium<ref>https://etcgroup.org/content/gene-drive-files</ref>. | ||
| Line 55: | Line 73: | ||
Because [[Target Malaria]] hopes to deploy their gene drives in [[African]] countries they have been at pains to emphasize independence from military agendas. However,Target Malaria’s [[Andrea Crisanti]] (working at [[Imperial College]]) is also a lead grantee or subcontractor for DARPA’s Safe Genes project, having confirmed he has been hired by DARPA on a $2.5m contract<ref>https://www.theguardian.com/science/2017/dec/04/us-military-agency-invests-100m-in-genetic-extinction-technologies</ref>. | Because [[Target Malaria]] hopes to deploy their gene drives in [[African]] countries they have been at pains to emphasize independence from military agendas. However,Target Malaria’s [[Andrea Crisanti]] (working at [[Imperial College]]) is also a lead grantee or subcontractor for DARPA’s Safe Genes project, having confirmed he has been hired by DARPA on a $2.5m contract<ref>https://www.theguardian.com/science/2017/dec/04/us-military-agency-invests-100m-in-genetic-extinction-technologies</ref>. | ||
| − | ==Imperial College London== | + | ===Imperial College London=== |
[[Imperial College London]] has been a pioneer in gene drive research, with DARPA funding<ref>https://www.theguardian.com/science/2017/dec/04/us-military-agency-invests-100m-in-genetic-extinction-technologies</ref>. In trials 2016-2018, scientists succeeded in destroying a population of mosquitoes in a lab by introducing a genetic mutation that spread through the population and eventually [[sterilized]] all of the mosquitoes. In previous experiments, mosquitoes had small random mutations that immunized them against the gene drive. The Imperial College scientists created a gene drive that did not fall prey to this type of resistance.<ref>https://www.statnews.com/2018/09/24/gene-drive-lab-mosquitos/</ref> | [[Imperial College London]] has been a pioneer in gene drive research, with DARPA funding<ref>https://www.theguardian.com/science/2017/dec/04/us-military-agency-invests-100m-in-genetic-extinction-technologies</ref>. In trials 2016-2018, scientists succeeded in destroying a population of mosquitoes in a lab by introducing a genetic mutation that spread through the population and eventually [[sterilized]] all of the mosquitoes. In previous experiments, mosquitoes had small random mutations that immunized them against the gene drive. The Imperial College scientists created a gene drive that did not fall prey to this type of resistance.<ref>https://www.statnews.com/2018/09/24/gene-drive-lab-mosquitos/</ref> | ||
== Spreading in the population == | == Spreading in the population == | ||
| − | Since it can never more than double in frequency with each generation, a gene drive introduced in a single individual typically requires dozens of generations to affect a substantial fraction of a population. Alternatively, releasing drive-containing organisms in sufficient numbers can affect the rest within a few generations; for instance, by introducing it in every thousandth individual, it takes only 12–15 generations to be present in all individuals.<ref> | + | Since it can never more than double in frequency with each generation, a gene drive introduced in a single individual typically requires dozens of generations to affect a substantial fraction of a population. Alternatively, releasing drive-containing organisms in sufficient numbers can affect the rest within a few generations; for instance, by introducing it in every thousandth individual, it takes only 12–15 generations to be present in all individuals.<ref>Unckless RL, Messer PW, Connallon T, Clark AG, "Modeling the Manipulation of Natural Populations by the Mutagenic Chain Reaction", Genetics, volume 201, issue 2, pages 425–31, October 2015, doi 10.1534/genetics.115.177592</ref> Whether a gene drive will ultimately become fixed in a population and at which speed depends on its effect on individuals fitness, on the rate of allele conversion, and on the population structure. In a well mixed population and with realistic allele conversion frequencies (≈90%), population genetics predicts that gene drives get fixed for selection coefficient smaller than 0.3;<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4596658</ref> in other words, gene drives can be used to spread modifications as long as reproductive success is not reduced by more than 30%. This is in contrast with normal genes, which can only spread across large populations if they increase fitness. |
| − | + | Because the strategy usually relies on the simultaneous presence of an unmodified and a gene drive allele in the same [[cell nucleus]], it had generally been assumed that a gene drive could only be engineered in sexually reproducing organisms, excluding [[bacteria]] and [[Virus|viruses]]. However, technological breakthroughs have enabled the design of a gene drive strategy that doesn’t involve sexual reproduction, but relies on [[Coinfection|co-infection]] of a given cell by a naturally occurring and an engineered virus. Upon co-infection, the unmodified genome is cut and repaired by homologous recombination, producing new gene drive viruses that can progressively replace the naturally occurring population. In [[cell culture]] experiments, it was shown that a viral gene drive can spread into the viral population and strongly reduce the infectivity of the virus, which opens novel therapeutic strategies against herpesviruses.<ref>https://www.nature.com/articles/s41467-020-18678-0</ref> | |
| − | Because the strategy usually relies on the simultaneous presence of an unmodified and a gene drive allele in the same [[cell nucleus]], it had generally been assumed that a gene drive could only be engineered in sexually reproducing organisms, excluding [[bacteria]] and [[Virus|viruses]]. However, | ||
| − | |||
| − | |||
== Technical limitations == | == Technical limitations == | ||
| − | Because gene drives propagate by replacing other alleles that contain a cutting site and the corresponding homologies, their application has been mostly limited to sexually reproducing species (because they are [[Ploidy|diploid]] or [[Polyploid#Plants|polyploid]] and alleles are mixed at each generation). As a side effect, inbreeding could in principle be an escape mechanism, but the extent to which this can happen in practice is difficult to evaluate.<ref> | + | Because gene drives propagate by replacing other alleles that contain a cutting site and the corresponding homologies, their application has been mostly limited to sexually reproducing species (because they are [[Ploidy|diploid]] or [[Polyploid#Plants|polyploid]] and alleles are mixed at each generation). As a side effect, inbreeding could in principle be an escape mechanism, but the extent to which this can happen in practice is difficult to evaluate.<ref>James J. Bull, 2016-04-02, "Lethal Gene Drive Selects Escape through Inbreeding" biorxiv 10.1101/046847</ref> |
| − | Due to the number of generations required for a gene drive to affect an entire population, the time to universality varies according to the reproductive cycle of each species: it may require under a year for some invertebrates, but centuries for organisms with years-long intervals between birth and [[sexual maturity]], such as humans.<ref> | + | Due to the number of generations required for a gene drive to affect an entire population, the time to universality varies according to the reproductive cycle of each species: it may require under a year for some invertebrates, but centuries for organisms with years-long intervals between birth and [[sexual maturity]], such as humans.<ref>Oye KA, Esvelt K, Appleton E, Catteruccia F, Church G, Kuiken T, Lightfoot SB, McNamara J, Smidler A, Collins JP, "Biotechnology. Regulating gene drives", Science, volume 345, issue 6197, pages 626–8, August 2014, doi 10.1126/science.1254287</ref> Hence this technology is of most use in fast-reproducing species. |
| − | + | Other problematic issues highlighted by researchers include:<ref>https://web.archive.org/web/20140730041832/http://www.synbioproject.org/process/assets/files/6685/synbio_res_agenda1.pdf |archive-date=2014-07-30 |url-status=dead </ref> | |
| − | |||
* Mutations: A mutation could happen mid-drive, which has the potential to allow unwanted traits to "ride along". | * Mutations: A mutation could happen mid-drive, which has the potential to allow unwanted traits to "ride along". | ||
* Escape: Cross-breeding or [[gene flow]] potentially allow a drive to move beyond its target population. | * Escape: Cross-breeding or [[gene flow]] potentially allow a drive to move beyond its target population. | ||
| − | * Ecological impacts: Even when new traits' direct impact on a target is understood, the drive may have side effects on the surroundings. | + | * Ecological impacts: Even when new traits' direct impact on a target is understood, the drive may have side effects on the surroundings.' |
| + | ===Bioethics concerns=== | ||
The [[Broad Institute]] of MIT and Harvard added gene drives to a list of uses of gene-editing technology it doesn't think companies should pursue.<ref>https://www.technologyreview.com/s/609619/farmers-seek-to-deploy-powerful-gene-drive/|access-date=2018-04-28|language=en</ref> | The [[Broad Institute]] of MIT and Harvard added gene drives to a list of uses of gene-editing technology it doesn't think companies should pursue.<ref>https://www.technologyreview.com/s/609619/farmers-seek-to-deploy-powerful-gene-drive/|access-date=2018-04-28|language=en</ref> | ||
| − | + | Gene drives affect all future generations and represent the possibility of a larger change in a living species than has been possible before.<ref>https://www.pbs.org/wgbh/nova/next/evolution/crispr-gene-drives/ "I don't care if it's a weed or a blight, people still are going to say this is way too massive a genetic engineering project," [bioethicist] Caplan says. "Secondly, it's altering things that are inherited, and that's always been a bright line for genetic engineering."</ref> | |
| − | Gene drives affect all future generations and represent the possibility of a larger change in a living species than has been possible before.<ref> | ||
| − | In December 2015, scientists of major world academies called for a moratorium on inheritable [[human genome]] edits that would affect the germline, including those related to [[#CRISPR/Cas9|CRISPR-Cas9]] technologies,<ref> | + | In December 2015, scientists of major world academies called for a moratorium on inheritable [[human genome]] edits that would affect the germline, including those related to [[#CRISPR/Cas9|CRISPR-Cas9]] technologies,<ref>https://www.nytimes.com/2015/12/04/science/crispr-cas9-human-genome-editing-moratorium.html</ref> but supported continued basic research and gene editing that would not affect future generations.<ref>https://www.newscientist.com/article/mg22830514-500-geneticists-vote-to-allow-gene-editing-of-human-embryos/</ref> In February 2016, British scientists were given permission by regulators to genetically modify [[human embryos]] by using CRISPR-Cas9 and related techniques on condition that the embryos were destroyed in seven days.<ref>https://www.bbc.co.uk/news/health-35459054</ref><ref>https://web.archive.org/web/20160201114343/http://bigstory.ap.org/article/fdda5bf9f0314b748c7438c9659da83a/britain-approves-controversial-gene-editing-technique |archive-date=1 February 2016</ref> In June 2016, the US [[National Academies of Sciences, Engineering, and Medicine]] released a report on their "Recommendations for Responsible Conduct" of gene drives.<ref>http://nas-sites.org/gene-drives/|access-date=June 9, 2016</ref> |
Models suggest that extinction-oriented gene drives will wipe out target species and that drives could reach populations beyond the target given minimal connectivity between them.<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6014726</ref> | Models suggest that extinction-oriented gene drives will wipe out target species and that drives could reach populations beyond the target given minimal connectivity between them.<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6014726</ref> | ||
[[Kevin M. Esvelt]] stated that an open conversation was needed around the safety of gene drives: "In our view, it is wise to assume that invasive and self-propagating gene drive systems are likely to spread to every population of the target species throughout the world. Accordingly, they should only be built to combat true plagues such as malaria, for which we have few adequate countermeasures and that offer a realistic path towards an international agreement to deploy among all affected nations.".<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5689824</ref> He moved to an open model for his own research on using gene drive to eradicate [[Lyme disease]] (itself a biological weapon released by accident) in [[Nantucket]] and [[Martha's Vineyard]].<ref>https://www.theatlantic.com/science/archive/2017/07/a-scientists-plan-to-protect-the-world-by-changing-how-science-is-done/532962/</ref> Esvelt and colleagues suggested that CRISPR could be used to save endangered wildlife from extinction. Esvelt later retracted his support for the idea, except for extremely hazardous populations such as malaria-carrying mosquitoes and isolated islands that would prevent the drive from spreading beyond the target area.<ref>https://www.nytimes.com/2017/11/16/science/gene-drives-crispr.html</ref> | [[Kevin M. Esvelt]] stated that an open conversation was needed around the safety of gene drives: "In our view, it is wise to assume that invasive and self-propagating gene drive systems are likely to spread to every population of the target species throughout the world. Accordingly, they should only be built to combat true plagues such as malaria, for which we have few adequate countermeasures and that offer a realistic path towards an international agreement to deploy among all affected nations.".<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5689824</ref> He moved to an open model for his own research on using gene drive to eradicate [[Lyme disease]] (itself a biological weapon released by accident) in [[Nantucket]] and [[Martha's Vineyard]].<ref>https://www.theatlantic.com/science/archive/2017/07/a-scientists-plan-to-protect-the-world-by-changing-how-science-is-done/532962/</ref> Esvelt and colleagues suggested that CRISPR could be used to save endangered wildlife from extinction. Esvelt later retracted his support for the idea, except for extremely hazardous populations such as malaria-carrying mosquitoes and isolated islands that would prevent the drive from spreading beyond the target area.<ref>https://www.nytimes.com/2017/11/16/science/gene-drives-crispr.html</ref> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
== Mechanism == | == Mechanism == | ||
| Line 104: | Line 111: | ||
As a result, the gene drive insertion in the genome will re-occur in each organism that inherits one copy of the modification and one copy of the wild-type gene. If the gene drive is already present in the egg cell (e.g. when received from one parent), all the gametes of the individual will carry the gene drive (instead of 50% in the case of a normal gene).<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217</ref> | As a result, the gene drive insertion in the genome will re-occur in each organism that inherits one copy of the modification and one copy of the wild-type gene. If the gene drive is already present in the egg cell (e.g. when received from one parent), all the gametes of the individual will carry the gene drive (instead of 50% in the case of a normal gene).<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217</ref> | ||
| − | == Control strategies == | + | === Control strategies === |
Scientists have designed multiple strategies to maintain control over gene drives. | Scientists have designed multiple strategies to maintain control over gene drives. | ||
| − | The drosophila drive requires at least thousands of insects for the drive to begin. A few individuals escaping the target region would be unlikely to spread the drive.<ref>https://www.technologyreview.com/s/603533/first-gene-drive-in-mammals-could-aid-vast-new-zealand-eradication-plan/ | + | The drosophila drive requires at least thousands of insects for the drive to begin. A few individuals escaping the target region would be unlikely to spread the drive.<ref>https://www.technologyreview.com/s/603533/first-gene-drive-in-mammals-could-aid-vast-new-zealand-eradication-plan/</ref> |
| − | In 2020 researchers reported the development of two active [[guide RNA]]-only elements that, according to their study, may enable halting or deleting gene drives introduced into populations in the wild with [[CRISPR-Cas9 gene editing]]. The paper's senior author cautions that the two neutralizing systems they demonstrated in cage trials "should not be used with a [[Security#Perceptions of security|false sense of security]] for field-implemented gene drives".<ref> | + | In 2020 researchers reported the development of two active [[guide RNA]]-only elements that, according to their study, may enable halting or deleting gene drives introduced into populations in the wild with [[CRISPR-Cas9 gene editing]]. The paper's senior author cautions that the two neutralizing systems they demonstrated in cage trials "should not be used with a [[Security#Perceptions of security|false sense of security]] for field-implemented gene drives".<ref>[https://phys.org/news/2020-09-biologists-genetic-neutralize-gene.html Biologists create new genetic systems to neutralize gene drives</ref><ref>[https://www.sciencedirect.com/science/article/abs/pii/S1097276520306110 Active Genetic Neutralizing Elements for Halting or Deleting Gene Drives]</ref> |
== CRISPR == | == CRISPR == | ||
| − | [[CRISPR]]<ref> | + | [[CRISPR]]<ref>https://doi.org/10.1126%2Fscience.341.6148.833</ref> is a DNA editing method that makes genetic engineering faster, easier, and more efficient.<ref>https://www.nytimes.com/2015/05/12/science/jennifer-doudna-crispr-cas9-genetic-engineering.html</ref> The approach involves expressing an [[RNA]]-guided [[endonuclease]] such as Cas9 along with guide RNAs directing it to a particular sequence to be edited. When the endonuclease cuts the target sequence, the cell repairs the damage by replacing the original sequence with homologous DNA. By introducing an additional template with appropriate homologues, an endonuclease can be used to delete, add or modify genes in an unprecedentedly simple manner. {{As of|2014, it had been tested in cells of 20 species, including humans.<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217</ref> In many of these species, the edits modified the organism's [[germline]], allowing them to be inherited. |
| − | In 2014 Esvelt and coworkers first suggested that CRISPR/Cas9 might be used to build endonuclease gene drives.<ref> | + | In 2014 Esvelt and coworkers first suggested that CRISPR/Cas9 might be used to build endonuclease gene drives.<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217</ref> In 2015 researchers published successful engineering of CRISPR-based gene drives in ''[[Saccharomyces cerevisiae#Model organism|Saccharomyces]]''<ref>https://doi.org/10.1101%2F013896</ref>'','' ''[[Drosophila#Laboratory-cultured animals|Drosophila]]'',<ref>>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4687737</ref> and [[mosquito]]es.<ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4679060</ref><ref>https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4913862</ref> All four studies demonstrated efficient inheritance distortion over successive generations, with one study demonstrating the spread of a gene drive into laboratory populations.<ref> name=":0" </ref> Drive-resistant alleles were expected to arise for each of the described gene drives, however this could be delayed or prevented by targeting highly conserved sites at which resistance is expected to have a severe fitness cost. |
Because of CRISPR's targeting flexibility, gene drives could theoretically be used to engineer almost any trait. Unlike previous designs, they could be tailored to block the evolution of drive resistance in the target population by targeting multiple sequences within appropriate genes. CRISPR could permit a variety of gene drive architectures intended to control rather than crash populations. | Because of CRISPR's targeting flexibility, gene drives could theoretically be used to engineer almost any trait. Unlike previous designs, they could be tailored to block the evolution of drive resistance in the target population by targeting multiple sequences within appropriate genes. CRISPR could permit a variety of gene drive architectures intended to control rather than crash populations. | ||
| Line 124: | Line 131: | ||
* introduce a genetic modification in wild populations. Gene drives constitute a major development that makes possible previously unattainable changes. | * introduce a genetic modification in wild populations. Gene drives constitute a major development that makes possible previously unattainable changes. | ||
| − | Because of their unprecedented potential risk, safeguard mechanisms have been proposed and tested.<ref> name=":3" </ref><ref> | + | Because of their unprecedented potential risk, safeguard mechanisms have been proposed and tested.<ref> name=":3" </ref><ref>DiCarlo JE, Chavez A, Dietz SL, Esvelt KM, Church GM, "Safeguarding CRISPR-Cas9 gene drives in yeast", Nature Biotechnology, volume 33, issue 12, pages 1250–1255, December 2015, doi 10.1038/nbt.3412</ref> |
=== Disease vector species === | === Disease vector species === | ||
| − | One possible application is to genetically modify [[mosquito]]es and other disease vectors so that they cannot transmit diseases such as [[malaria]] and [[dengue fever]]. Researchers have claimed that by applying the technique to 1% of the wild population of mosquitoes, that they could eradicate malaria within a year.<ref> | + | One possible application is to genetically modify [[mosquito]]es and other disease vectors so that they cannot transmit diseases such as [[malaria]] and [[dengue fever]]. Researchers have claimed that by applying the technique to 1% of the wild population of mosquitoes, that they could eradicate malaria within a year.<ref>[https://www.youtube.com/watch?v=OI_OhvOumT0 Gene editing can now change an entire species — forever], 2016-06-02, [[Jennifer Kahn]], [[TED]]</ref> |
=== Invasive species control === | === Invasive species control === | ||
| − | A gene drive could be used to eliminate [[invasive species]] and has, for example, been proposed as a way to eliminate [[invasive species in New Zealand]].<ref> | + | A gene drive could be used to eliminate [[invasive species]] and has, for example, been proposed as a way to eliminate [[invasive species in New Zealand]].<ref>http://www.merlinnz.com/blog/crispr-pest-free-nz/</ref> Gene drives for biodiversity conservation purposes are being explored as part of The Genetic Biocontrol of Invasive Rodents (GBIRd) program because they offer the potential for reduced risk to non-target species and reduced costs when compared to traditional invasive species removal techniques. Given the risks of such an approach described below, the GBIRd partnership is committed to a deliberate, step-wise process that will only proceed with public alignment, as recommended by the world's leading gene drive researchers from the Australian and US National Academy of |
Sciences and many others.<ref>http://www.geneticbiocontrol.org/wp-content/uploads/2018/05/GBIRD-FactSheet-April-2018.pdf </ref> A wider Outreach Network for Gene Drive Research exists to raise awareness of the value of gene drive research for the public good.<ref>https://genedrivenetwork.org/resources/6-mission-principles-statement-july2018/file </ref> | Sciences and many others.<ref>http://www.geneticbiocontrol.org/wp-content/uploads/2018/05/GBIRD-FactSheet-April-2018.pdf </ref> A wider Outreach Network for Gene Drive Research exists to raise awareness of the value of gene drive research for the public good.<ref>https://genedrivenetwork.org/resources/6-mission-principles-statement-july2018/file </ref> | ||
| − | Some scientists are concerned about the technique, fearing it could spread and wipe out species in native habitats.<ref> | + | Some scientists are concerned about the technique, fearing it could spread and wipe out species in native habitats.<ref>http://theconversation.com/gene-drives-could-wipe-out-whole-populations-of-pests-in-one-fell-swoop-81681 </ref> The gene could mutate, potentially causing unforeseen problems (as could any gene).<ref>http://blogs.plos.org/dnascience/2017/11/30/an-argument-against-gene-drives-to-extinguish-new-zealand-mammals-life-finds-a-way/</ref> Many non-native species can hybridize with native species, such that a gene drive afflicting a non-native plant or animal that hybridizes with a native species could doom the native species. Many non-native species have naturalized into their new environment so well that crops and/or native species have adapted to depend on them.<ref>https://www.odt.co.nz/opinion/risks-may-accompany-gene-drive-technology#comment-1086</ref> |
==== New Zealand ==== | ==== New Zealand ==== | ||
| − | + | The Predator Free 2050 project is a New Zealand government program to completely eliminate eight invasive mammalian predator species (including rats, short-tailed weasels, and possums) from the country by 2050.<ref>https://www.wired.com/2016/07/new-zealand-plans-kill-non-human-invasive-mammals/</ref><ref>http://www.scoop.co.nz/stories/SC1701/S00024/predator-free-nz-expert-qa.htm</ref> The projects was first announced in 2016 by New Zealand's prime minister [[John Key]] and in January 2017 it was announced that gene drives would be used in the effort.<ref>http://www.scoop.co.nz/stories/SC1701/S00024/predator-free-nz-expert-qa.htm</ref> In 2017 one group in Australia and another in Texas released preliminary research into creating 'daughterless mice', using gene drives in mammals.<ref>https://www.technologyreview.com/s/603533/first-gene-drive-in-mammals-could-aid-vast-new-zealand-eradication-plan/</ref> | |
| − | The Predator Free 2050 project is a New Zealand government program to completely eliminate eight invasive mammalian predator species (including rats, short-tailed weasels, and possums) from the country by 2050.<ref>https://www.wired.com/2016/07/new-zealand-plans-kill-non-human-invasive-mammals/</ref><ref> | ||
==== California ==== | ==== California ==== | ||
| − | In 2017 scientists at the University of California, Riverside developed a gene drive to attack the [[Invasive species|invasive]] [[Drosophila suzukii|spotted-wing drosophila]], a type of fruit fly native to Asia that is costing California's cherry farms $700 million per year because of its tail's razor-edged “[[ovipositor]]” that destroys unblemished fruit. The primary alternative control strategy involves the use of [[insecticide]]s called [[pyrethroid]]s that kills almost all insects that it contacts.<ref>https://www.technologyreview.com/s/609619/farmers-seek-to-deploy-powerful-gene-drive/</ref> | + | In 2017 scientists at the University of California, Riverside developed a gene drive to attack the [[Invasive species|invasive]] [[Drosophila suzukii|spotted-wing drosophila]], a type of fruit fly native to Asia that is costing [[California]]'s cherry farms $700 million per year because of its tail's razor-edged “[[ovipositor]]” that destroys unblemished fruit. The primary alternative control strategy involves the use of [[insecticide]]s called [[pyrethroid]]s that kills almost all insects that it contacts.<ref>https://www.technologyreview.com/s/609619/farmers-seek-to-deploy-powerful-gene-drive/</ref> |
=== Wild animal welfare === | === Wild animal welfare === | ||
| − | The transhumanist philosopher [[David Pearce (philosopher)|David Pearce]] has advocated for using CRISPR-based gene drives to reduce the [[suffering of wild animals]].<ref>>https://digitalcommons.calpoly.edu/bts/vol23/iss1/8</ref> [[Kevin M. Esvelt]], an American biologist who has helped develop gene drive technology, has argued that there is a moral case for the elimination of the [[New World screwworm]] through such technologies because of the immense suffering that infested wild animals experience when they are eaten alive.<ref>https://leapsmag.com/when-are-we-obligated-to-edit-wild-creatures/</ref> | + | The [[transhumanist]] philosopher [[David Pearce (philosopher)|David Pearce]] has advocated for using CRISPR-based gene drives to reduce the [[suffering of wild animals]].<ref>>https://digitalcommons.calpoly.edu/bts/vol23/iss1/8</ref> [[Kevin M. Esvelt]], an American biologist who has helped develop gene drive technology, has argued that there is a moral case for the elimination of the [[New World screwworm]] through such technologies because of the immense suffering that infested wild animals experience when they are eaten alive.<ref>https://leapsmag.com/when-are-we-obligated-to-edit-wild-creatures/</ref> |
== External links == | == External links == | ||
* [https://genedrivenetwork.org The Outreach Network for Gene Drive Research website] | * [https://genedrivenetwork.org The Outreach Network for Gene Drive Research website] | ||
* [http://www.geneticbiocontrol.org/ The Genetic Biocontrol of Invasive Rodents (GBIRd) program website] | * [http://www.geneticbiocontrol.org/ The Genetic Biocontrol of Invasive Rodents (GBIRd) program website] | ||
| − | * | + | * [https://www.youtube.com/watch?v=Cy69C6vnFCQ/ Gene Drive from Harvard's Wyss Institute]] |
{{SMWDocs}} | {{SMWDocs}} | ||
==References== | ==References== | ||
<references/> | <references/> | ||
Latest revision as of 20:00, 1 February 2022
(Biological weapon/Research, research, Genetic engineering) | |
|---|---|
| Interest of | Bill Gates |
A gene drive is an existing technology of genetic engineering that is able to propagate a particular suite of genes throughout a population[1] by altering the probability that a specific allele will be transmitted to offspring (instead of the Mendelian 50% probability). It application is particularly suited for creating an irreversible species extinction[2].
The U.S. Defense Advanced Research Projects Agency (DARPA) has given approximately $100 million for gene drive research, making them likely the largest single funder of gene drive research on the planet. The secretive top-level JASON group of military advisors produced a classified study on gene drive in 2017, reflecting an extremely high level of interest and activity by other sections of the U.S. military and Intelligence community[3].
The Bill and Melinda Gates Foundation paid a PR firm $1.6 million to secretly stack key UN advisory processes with gene drive-friendly scientists[4].
Contents
A Powerful Technique
Gene drives can arise through a variety of mechanisms.[5][6] They have been proposed to provide an effective means of genetically modifying specific populations and entire species.
The technique can employ adding, deleting, disrupting, or modifying genes.[7][8]
Proposed applications include exterminating insects that carry pathogens (notably mosquitoes that transmit malaria, dengue, and zika pathogens), controlling invasive species, or eliminating herbicide or pesticide resistance.[9][10][11][12]
As with any potentially powerful technique, gene drives can be misused in a variety of ways or induce unintended consequences. For example, a gene drive intended to affect only a local population might spread across an entire species. Gene drives used to eradicate populations of invasive species in their non-native habitats may have consequences for the population of the species as a whole, even in its native habitat. Any accidental return of individuals of the species to its original habitats, through natural migration, environmental disruption (storms, floods, etc.), accidental human transportation, or purposeful relocation, could unintentionally drive the species to extinction if the relocated individuals carried harmful gene drives.[13]
“It is very much easier to kill or sterilise a plant using gene editing than it is to make it herbicide or insect-resistant.”
Guy Reeves, expert in GM insects at the Max Planck Institute for Evolutionary Biology (2018) [14]
Gene drives can be built from many naturally occurring selfish genetic elements that use a variety of molecular mechanisms.[15] These naturally occurring mechanisms induce similar segregation distortion in the wild, arising when alleles evolve molecular mechanisms that give them a transmission chance greater than the normal 50%.
Most gene drives have been developed in insects, notably mosquitoes, as a way to control insect-borne pathogens. Recent developments designed gene drives directly in viruses, notably herpesviruses. These viral gene drives can propagate a modification into the population of viruses, and aim to reduce the infectivity of the virus.[16][17]
History
Austin Burt, an evolutionary geneticist at Imperial College London, introduced the possibility of conducting gene drives based on natural homing endonuclease selfish genetic elements in 2003.[18]
Researchers had already shown that such genes could act selfishly to spread rapidly over successive generations. Burt suggested that gene drives might be used to prevent a mosquito population from transmitting the malaria parasite or to crash a mosquito population. Gene drives based on homing endonucleases have been demonstrated in the laboratory in transgenic populations of mosquitoes[19] and fruit flies.[20][21] However, homing endonucleases are sequence-specific. Altering their specificity to target other sequences of interest remains a major challenge.[22] The possible applications of gene drive remained limited until the discovery of CRISPR and associated RNA-guided endonucleases such as Cas9 and Cpf1.
In June 2014, the World Health Organization (WHO) Special Programme for Research and Training in Tropical Diseases[23] issued guidelines[24] for evaluating genetically modified mosquitoes. In 2013 the European Food Safety Authority issued a protocol[25] for environmental assessments of all genetically modified organisms.
Interested parties
Defense Advanced Research Projects Agency
In December 2017, documents released under the US Freedom of Information Act showed that the military research agency DARPA had invested $100 million in gene drive research,[26], making them likely the largest single funder of gene drive research on the planet. The documents also reveal that DARPA either funds or co-ordinates with almost all major players working on gene drive development as well as the key holders of patents on CRISPR gene editing technology.
“Given that Darpa is a military agency, we find it surprising that the obvious and concerning dual-use aspects of this research have received so little attention,”
Felix Beck - lawyer at the University of Freiburg (2018) [14]
JASON Military Advisory Group
The secretive top-level JASON group of military advisors produced a classified study on gene drive in 2017, reflecting an extremely high level of interest and activity by other sections of the U.S. military and Intelligence community[27].
Emails show that the JASON study was initiated with a two day meeting of a select group of invited gene drive researchers in June 2017. At the meeting, the VP of Global Biotechnology for Monsanto gave a presentation on crop science and gene drives[28].
The Intelligence Advanced Research Projects Agency
The Intelligence Advanced Research Projects Agency (IARPA), an organization within the Office of the Director of National Intelligence, has also expressed interest in funding gene drive work. A scientist involved describes IARPA as “basically the intelligence agencies version of DARPA, which may be more frightening"[29]
Project for the New American Century
In their 2000 paper: "Rebuilding America's Defences: Strategies, Forces And Resources For A New Century", PNAC commented on biowarfare application of a then unnamed technology:
“And advanced forms of biological warfare that can “target” specific genotypes may transform biological warfare from the realm of terror to a politically useful tool.”
Project for the New American Century [30]
Bill & Melinda Gates Foundation
The Bill and Melinda Gates Foundation paid a PR firm $1.6 million to secretly stack key UN advisory processes with gene drive-friendly scientists, and recruited ostensibly independent academics and public officials into a private collaboration to counteract proposed regulations and to resist calls by scientists and conservationists for an international moratorium[31].
Target Malaria, a project funded by the Bill and Melinda Gates Foundation, invested $75 million in gene drive technology. The foundation originally estimated the technology to be ready for field use by 2029 somewhere in Africa. However, in 2016 Gates changed this estimate to some time within the following two years.[32]
Because Target Malaria hopes to deploy their gene drives in African countries they have been at pains to emphasize independence from military agendas. However,Target Malaria’s Andrea Crisanti (working at Imperial College) is also a lead grantee or subcontractor for DARPA’s Safe Genes project, having confirmed he has been hired by DARPA on a $2.5m contract[33].
Imperial College London
Imperial College London has been a pioneer in gene drive research, with DARPA funding[34]. In trials 2016-2018, scientists succeeded in destroying a population of mosquitoes in a lab by introducing a genetic mutation that spread through the population and eventually sterilized all of the mosquitoes. In previous experiments, mosquitoes had small random mutations that immunized them against the gene drive. The Imperial College scientists created a gene drive that did not fall prey to this type of resistance.[35]
Spreading in the population
Since it can never more than double in frequency with each generation, a gene drive introduced in a single individual typically requires dozens of generations to affect a substantial fraction of a population. Alternatively, releasing drive-containing organisms in sufficient numbers can affect the rest within a few generations; for instance, by introducing it in every thousandth individual, it takes only 12–15 generations to be present in all individuals.[36] Whether a gene drive will ultimately become fixed in a population and at which speed depends on its effect on individuals fitness, on the rate of allele conversion, and on the population structure. In a well mixed population and with realistic allele conversion frequencies (≈90%), population genetics predicts that gene drives get fixed for selection coefficient smaller than 0.3;[37] in other words, gene drives can be used to spread modifications as long as reproductive success is not reduced by more than 30%. This is in contrast with normal genes, which can only spread across large populations if they increase fitness.
Because the strategy usually relies on the simultaneous presence of an unmodified and a gene drive allele in the same cell nucleus, it had generally been assumed that a gene drive could only be engineered in sexually reproducing organisms, excluding bacteria and viruses. However, technological breakthroughs have enabled the design of a gene drive strategy that doesn’t involve sexual reproduction, but relies on co-infection of a given cell by a naturally occurring and an engineered virus. Upon co-infection, the unmodified genome is cut and repaired by homologous recombination, producing new gene drive viruses that can progressively replace the naturally occurring population. In cell culture experiments, it was shown that a viral gene drive can spread into the viral population and strongly reduce the infectivity of the virus, which opens novel therapeutic strategies against herpesviruses.[38]
Technical limitations
Because gene drives propagate by replacing other alleles that contain a cutting site and the corresponding homologies, their application has been mostly limited to sexually reproducing species (because they are diploid or polyploid and alleles are mixed at each generation). As a side effect, inbreeding could in principle be an escape mechanism, but the extent to which this can happen in practice is difficult to evaluate.[39]
Due to the number of generations required for a gene drive to affect an entire population, the time to universality varies according to the reproductive cycle of each species: it may require under a year for some invertebrates, but centuries for organisms with years-long intervals between birth and sexual maturity, such as humans.[40] Hence this technology is of most use in fast-reproducing species.
Other problematic issues highlighted by researchers include:[41]
- Mutations: A mutation could happen mid-drive, which has the potential to allow unwanted traits to "ride along".
- Escape: Cross-breeding or gene flow potentially allow a drive to move beyond its target population.
- Ecological impacts: Even when new traits' direct impact on a target is understood, the drive may have side effects on the surroundings.'
Bioethics concerns
The Broad Institute of MIT and Harvard added gene drives to a list of uses of gene-editing technology it doesn't think companies should pursue.[42]
Gene drives affect all future generations and represent the possibility of a larger change in a living species than has been possible before.[43]
In December 2015, scientists of major world academies called for a moratorium on inheritable human genome edits that would affect the germline, including those related to CRISPR-Cas9 technologies,[44] but supported continued basic research and gene editing that would not affect future generations.[45] In February 2016, British scientists were given permission by regulators to genetically modify human embryos by using CRISPR-Cas9 and related techniques on condition that the embryos were destroyed in seven days.[46][47] In June 2016, the US National Academies of Sciences, Engineering, and Medicine released a report on their "Recommendations for Responsible Conduct" of gene drives.[48]
Models suggest that extinction-oriented gene drives will wipe out target species and that drives could reach populations beyond the target given minimal connectivity between them.[49]
Kevin M. Esvelt stated that an open conversation was needed around the safety of gene drives: "In our view, it is wise to assume that invasive and self-propagating gene drive systems are likely to spread to every population of the target species throughout the world. Accordingly, they should only be built to combat true plagues such as malaria, for which we have few adequate countermeasures and that offer a realistic path towards an international agreement to deploy among all affected nations.".[50] He moved to an open model for his own research on using gene drive to eradicate Lyme disease (itself a biological weapon released by accident) in Nantucket and Martha's Vineyard.[51] Esvelt and colleagues suggested that CRISPR could be used to save endangered wildlife from extinction. Esvelt later retracted his support for the idea, except for extremely hazardous populations such as malaria-carrying mosquitoes and isolated islands that would prevent the drive from spreading beyond the target area.[52]
Mechanism
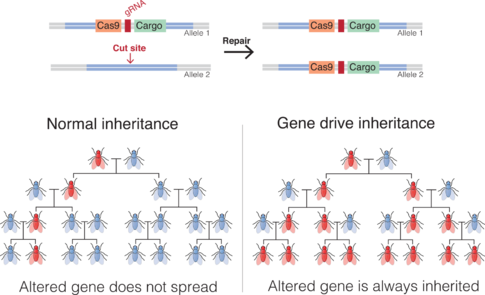
In sexually-reproducing species, most genes are present in two copies (which can be the same or different alleles), either one of which has a 50% chance of passing to a descendant. By biasing the inheritance of particular altered genes, synthetic gene drives could spread alterations through a population.[53][54]
Molecular mechanisms
At the molecular level, an endonuclease gene drive works by cutting a chromosome at a specific site that does not encode the drive, inducing the cell to repair the damage by copying the drive sequence onto the damaged chromosome. The cell then has two copies of the drive sequence. The method derives from genome editing techniques.
As a result, the gene drive insertion in the genome will re-occur in each organism that inherits one copy of the modification and one copy of the wild-type gene. If the gene drive is already present in the egg cell (e.g. when received from one parent), all the gametes of the individual will carry the gene drive (instead of 50% in the case of a normal gene).[55]
Control strategies
Scientists have designed multiple strategies to maintain control over gene drives.
The drosophila drive requires at least thousands of insects for the drive to begin. A few individuals escaping the target region would be unlikely to spread the drive.[56]
In 2020 researchers reported the development of two active guide RNA-only elements that, according to their study, may enable halting or deleting gene drives introduced into populations in the wild with CRISPR-Cas9 gene editing. The paper's senior author cautions that the two neutralizing systems they demonstrated in cage trials "should not be used with a false sense of security for field-implemented gene drives".[57][58]
CRISPR
CRISPR[59] is a DNA editing method that makes genetic engineering faster, easier, and more efficient.[60] The approach involves expressing an RNA-guided endonuclease such as Cas9 along with guide RNAs directing it to a particular sequence to be edited. When the endonuclease cuts the target sequence, the cell repairs the damage by replacing the original sequence with homologous DNA. By introducing an additional template with appropriate homologues, an endonuclease can be used to delete, add or modify genes in an unprecedentedly simple manner. {{As of|2014, it had been tested in cells of 20 species, including humans.[61] In many of these species, the edits modified the organism's germline, allowing them to be inherited.
In 2014 Esvelt and coworkers first suggested that CRISPR/Cas9 might be used to build endonuclease gene drives.[62] In 2015 researchers published successful engineering of CRISPR-based gene drives in Saccharomyces[63], Drosophila,[64] and mosquitoes.[65][66] All four studies demonstrated efficient inheritance distortion over successive generations, with one study demonstrating the spread of a gene drive into laboratory populations.[67] Drive-resistant alleles were expected to arise for each of the described gene drives, however this could be delayed or prevented by targeting highly conserved sites at which resistance is expected to have a severe fitness cost.
Because of CRISPR's targeting flexibility, gene drives could theoretically be used to engineer almost any trait. Unlike previous designs, they could be tailored to block the evolution of drive resistance in the target population by targeting multiple sequences within appropriate genes. CRISPR could permit a variety of gene drive architectures intended to control rather than crash populations.
Applications
Gene drives have two main classes of application, which have implications of different significance:
- introduce a genetic modification in laboratory populations; once a strain or a line carrying the gene drive has been produced, the drive can be passed to any other line by mating. Here the gene drive is used to achieve much more easily a task that could be accomplished with other techniques.
- introduce a genetic modification in wild populations. Gene drives constitute a major development that makes possible previously unattainable changes.
Because of their unprecedented potential risk, safeguard mechanisms have been proposed and tested.[68][69]
Disease vector species
One possible application is to genetically modify mosquitoes and other disease vectors so that they cannot transmit diseases such as malaria and dengue fever. Researchers have claimed that by applying the technique to 1% of the wild population of mosquitoes, that they could eradicate malaria within a year.[70]
Invasive species control
A gene drive could be used to eliminate invasive species and has, for example, been proposed as a way to eliminate invasive species in New Zealand.[71] Gene drives for biodiversity conservation purposes are being explored as part of The Genetic Biocontrol of Invasive Rodents (GBIRd) program because they offer the potential for reduced risk to non-target species and reduced costs when compared to traditional invasive species removal techniques. Given the risks of such an approach described below, the GBIRd partnership is committed to a deliberate, step-wise process that will only proceed with public alignment, as recommended by the world's leading gene drive researchers from the Australian and US National Academy of Sciences and many others.[72] A wider Outreach Network for Gene Drive Research exists to raise awareness of the value of gene drive research for the public good.[73]
Some scientists are concerned about the technique, fearing it could spread and wipe out species in native habitats.[74] The gene could mutate, potentially causing unforeseen problems (as could any gene).[75] Many non-native species can hybridize with native species, such that a gene drive afflicting a non-native plant or animal that hybridizes with a native species could doom the native species. Many non-native species have naturalized into their new environment so well that crops and/or native species have adapted to depend on them.[76]
New Zealand
The Predator Free 2050 project is a New Zealand government program to completely eliminate eight invasive mammalian predator species (including rats, short-tailed weasels, and possums) from the country by 2050.[77][78] The projects was first announced in 2016 by New Zealand's prime minister John Key and in January 2017 it was announced that gene drives would be used in the effort.[79] In 2017 one group in Australia and another in Texas released preliminary research into creating 'daughterless mice', using gene drives in mammals.[80]
California
In 2017 scientists at the University of California, Riverside developed a gene drive to attack the invasive spotted-wing drosophila, a type of fruit fly native to Asia that is costing California's cherry farms $700 million per year because of its tail's razor-edged “ovipositor” that destroys unblemished fruit. The primary alternative control strategy involves the use of insecticides called pyrethroids that kills almost all insects that it contacts.[81]
Wild animal welfare
The transhumanist philosopher David Pearce has advocated for using CRISPR-based gene drives to reduce the suffering of wild animals.[82] Kevin M. Esvelt, an American biologist who has helped develop gene drive technology, has argued that there is a moral case for the elimination of the New World screwworm through such technologies because of the immense suffering that infested wild animals experience when they are eaten alive.[83]
External links
- The Outreach Network for Gene Drive Research website
- The Genetic Biocontrol of Invasive Rodents (GBIRd) program website
- Gene Drive from Harvard's Wyss Institute]
Related Quotations
| Page | Quote | Author | Date |
|---|---|---|---|
| Biological weapon | “Fabricating scary narratives about superbugs is much easier than delivering on promises of making those bugs in labs. This is also a very productive avenue as people are woefully gullible and thus can be controlled by narratives just as effectively as by an actual scary-scary bioengineered virus. Biodefense is a huge grift on both sides. 'Their' side appropriates money and power, and new billion dollar agencies for 'Pandemic Preparedness'. 'Our' side gets millions of followers talking about them evil guys, or spinning stories about biolabs in Wuhan, Ukraine and lately California. They leak' from labs almost every week, and using CRISPER gene drive narrative logic, all mice in the world should look like Ralph Baric by now.” | Sasha Latypova | November 2023 |
| Transhumanism | “Even if half the world’s species were lost [during genetic experiments], enormous diversity would still remain. When those in the distant future look back on this period of history, they will likely see it not as the era when the natural environment was impoverished, but as the age when a plethora of new forms—some biological, some technological, some a combination of the two—burst onto the scene. We best serve ourselves, as well as future generations, by focusing on the short-term consequences of our actions rather than our vague notions about the needs of the distant future.” | Gregory Stock | 1993 |
Sponsor
| Event | Description |
|---|---|
| Open Philanthropy | Grant maker funneling deep state money among other things to pandemic planning. Financed Event 201. |
References
- ↑ >https://www.nature.com/news/us-defence-agencies-grapple-with-gene-drives-1.22345
- ↑ https://geneticliteracyproject.org/2020/08/18/viewpoint-is-there-a-scientific-basis-to-ban-gene-drive-technology-that-can-rid-us-of-virus-carrying-rodents-and-mosquitoes/
- ↑ http://genedrivefiles.synbiowatch.org/2017/12/01/us-military-gene-drive-development/
- ↑ https://etcgroup.org/content/gene-drive-files
- ↑ https://doi.org/10.1038%2Fnrg.2015.34
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6195636
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1691325
- ↑ http://news.sciencemag.org/biology/2014/07/u-s-researchers-call-greater-oversight-powerful-genetic-technology%7Caccess-date=2014-07-18
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217
- ↑ https://doi.org/10.1089%2Fvbz.2007.0273
- ↑ https://portals.iucn.org/library/node/48408
- ↑ https://www.wired.com/story/this-gene-editing-tech-might-be-too-dangerous-to-unleash/
- ↑ a b https://www.independent.co.uk/news/science/us-military-plan-biological-weapons-insect-allies-virus-crop-darpa-a8568996.html
- ↑ Champer J, Buchman A, Akbari OS, "Cheating evolution: engineering gene drives to manipulate the fate of wild populations", Nature Reviews. Genetics, volume 17, issue 3, pages 146–59, March 2016, doi = 10.1038/nrg.2015.34
- ↑ https://www.nature.com/articles/s41467-020-18678-0
- ↑ https://www.acsh.org/news/2020/09/30/gene-drives-could-kill-mosquitoes-and-suppress-herpesvirus-infections-15060
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1691325
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3093433
- ↑ >https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3120159
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3548849
- ↑ https://doi.org/10.1038%2Fnrg.2015.34
- ↑ https://www.who.int/tdr/about/en/%7Caccess-date=2014-07-18
- ↑ https://www.who.int/tdr/news/2014/framwk-eval-gm-mosquitoes/en%7Caccess-date=2014-07-18
- ↑ http://www.efsa.europa.eu/en/efsajournal/pub/3200
- ↑ https://www.theguardian.com/science/2017/dec/04/us-military-agency-invests-100m-in-genetic-extinction-technologies
- ↑ http://genedrivefiles.synbiowatch.org/2017/12/01/us-military-gene-drive-development/
- ↑ http://genedrivefiles.synbiowatch.org/2017/12/01/us-military-gene-drive-development/
- ↑ http://genedrivefiles.synbiowatch.org/20170716-fw_-gbird-update-engagements-and-other-important-items-472/
- ↑ https://web.archive.org/web/20020923154604/http://www.newamericancentury.org/RebuildingAmericasDefenses.pdf
- ↑ https://etcgroup.org/content/gene-drive-files
- ↑ https://www.technologyreview.com/s/602304/bill-gates-doubles-his-bet-on-wiping-out-mosquitoes-with-gene-editing/
- ↑ https://www.theguardian.com/science/2017/dec/04/us-military-agency-invests-100m-in-genetic-extinction-technologies
- ↑ https://www.theguardian.com/science/2017/dec/04/us-military-agency-invests-100m-in-genetic-extinction-technologies
- ↑ https://www.statnews.com/2018/09/24/gene-drive-lab-mosquitos/
- ↑ Unckless RL, Messer PW, Connallon T, Clark AG, "Modeling the Manipulation of Natural Populations by the Mutagenic Chain Reaction", Genetics, volume 201, issue 2, pages 425–31, October 2015, doi 10.1534/genetics.115.177592
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4596658
- ↑ https://www.nature.com/articles/s41467-020-18678-0
- ↑ James J. Bull, 2016-04-02, "Lethal Gene Drive Selects Escape through Inbreeding" biorxiv 10.1101/046847
- ↑ Oye KA, Esvelt K, Appleton E, Catteruccia F, Church G, Kuiken T, Lightfoot SB, McNamara J, Smidler A, Collins JP, "Biotechnology. Regulating gene drives", Science, volume 345, issue 6197, pages 626–8, August 2014, doi 10.1126/science.1254287
- ↑ https://web.archive.org/web/20140730041832/http://www.synbioproject.org/process/assets/files/6685/synbio_res_agenda1.pdf |archive-date=2014-07-30 |url-status=dead
- ↑ https://www.technologyreview.com/s/609619/farmers-seek-to-deploy-powerful-gene-drive/%7Caccess-date=2018-04-28%7Clanguage=en
- ↑ https://www.pbs.org/wgbh/nova/next/evolution/crispr-gene-drives/ "I don't care if it's a weed or a blight, people still are going to say this is way too massive a genetic engineering project," [bioethicist] Caplan says. "Secondly, it's altering things that are inherited, and that's always been a bright line for genetic engineering."
- ↑ https://www.nytimes.com/2015/12/04/science/crispr-cas9-human-genome-editing-moratorium.html
- ↑ https://www.newscientist.com/article/mg22830514-500-geneticists-vote-to-allow-gene-editing-of-human-embryos/
- ↑ https://www.bbc.co.uk/news/health-35459054
- ↑ https://web.archive.org/web/20160201114343/http://bigstory.ap.org/article/fdda5bf9f0314b748c7438c9659da83a/britain-approves-controversial-gene-editing-technique |archive-date=1 February 2016
- ↑ http://nas-sites.org/gene-drives/%7Caccess-date=June 9, 2016
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6014726
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5689824
- ↑ https://www.theatlantic.com/science/archive/2017/07/a-scientists-plan-to-protect-the-world-by-changing-how-science-is-done/532962/
- ↑ https://www.nytimes.com/2017/11/16/science/gene-drives-crispr.html
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1691325
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217
- ↑ https://www.technologyreview.com/s/603533/first-gene-drive-in-mammals-could-aid-vast-new-zealand-eradication-plan/
- ↑ [https://phys.org/news/2020-09-biologists-genetic-neutralize-gene.html Biologists create new genetic systems to neutralize gene drives
- ↑ Active Genetic Neutralizing Elements for Halting or Deleting Gene Drives
- ↑ https://doi.org/10.1126%2Fscience.341.6148.833
- ↑ https://www.nytimes.com/2015/05/12/science/jennifer-doudna-crispr-cas9-genetic-engineering.html
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4117217
- ↑ https://doi.org/10.1101%2F013896
- ↑ >https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4687737
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4679060
- ↑ https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4913862
- ↑ name=":0"
- ↑ name=":3"
- ↑ DiCarlo JE, Chavez A, Dietz SL, Esvelt KM, Church GM, "Safeguarding CRISPR-Cas9 gene drives in yeast", Nature Biotechnology, volume 33, issue 12, pages 1250–1255, December 2015, doi 10.1038/nbt.3412
- ↑ Gene editing can now change an entire species — forever, 2016-06-02, Jennifer Kahn, TED
- ↑ http://www.merlinnz.com/blog/crispr-pest-free-nz/
- ↑ http://www.geneticbiocontrol.org/wp-content/uploads/2018/05/GBIRD-FactSheet-April-2018.pdf
- ↑ https://genedrivenetwork.org/resources/6-mission-principles-statement-july2018/file
- ↑ http://theconversation.com/gene-drives-could-wipe-out-whole-populations-of-pests-in-one-fell-swoop-81681
- ↑ http://blogs.plos.org/dnascience/2017/11/30/an-argument-against-gene-drives-to-extinguish-new-zealand-mammals-life-finds-a-way/
- ↑ https://www.odt.co.nz/opinion/risks-may-accompany-gene-drive-technology#comment-1086
- ↑ https://www.wired.com/2016/07/new-zealand-plans-kill-non-human-invasive-mammals/
- ↑ http://www.scoop.co.nz/stories/SC1701/S00024/predator-free-nz-expert-qa.htm
- ↑ http://www.scoop.co.nz/stories/SC1701/S00024/predator-free-nz-expert-qa.htm
- ↑ https://www.technologyreview.com/s/603533/first-gene-drive-in-mammals-could-aid-vast-new-zealand-eradication-plan/
- ↑ https://www.technologyreview.com/s/609619/farmers-seek-to-deploy-powerful-gene-drive/
- ↑ >https://digitalcommons.calpoly.edu/bts/vol23/iss1/8
- ↑ https://leapsmag.com/when-are-we-obligated-to-edit-wild-creatures/